Vascular endothelial growth factor (VEGF)-related single nucleotide polymorphisms rs10738760 and rs6921438 are not associated with diabetic retinopathy (DR) in Slovenian patients with type 2 diabetes mellitus (T2DM)
DOI:
https://doi.org/10.17305/bjbms.2017.2068Keywords:
Vascular endothelial growth factor, VEGF, VEGFA, diabetic retinopathy, DR, type 2 diabetes mellitus, T2DM, single nucleotide polymorphism, SNPAbstract
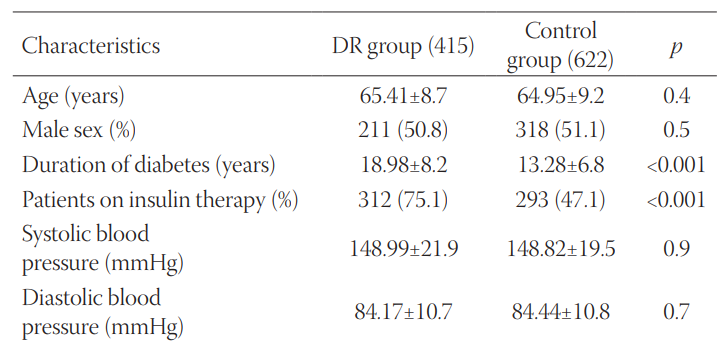
Diabetic retinopathy (DR) is a complication of diabetes characterized by vascular permeability, increased tissue ischemia, and angiogenesis. One of the most important proteins involved in angiogenesis is vascular endothelial growth factor (VEGF, also known as VEGFA). A previous study demonstrated that two single nucleotide polymorphisms (SNPs), rs6921438 and rs10738760, account for nearly half the variation in circulating VEGF levels. The aim of our study was to assess the association between rs6921438 and rs10738760 and DR in Slovenian patients with type 2 diabetes mellitus (T2DM). This case-control study enrolled 1037 unrelated Slovenian individuals (Caucasians) with T2DM. DR group included 415 T2DM patients with DR, while control group included 622 T2DM patients with no clinical signs of DR. The clinical and laboratory data were obtained from the medical records of the patients. The genotyping of rs6921438 and rs10738760 SNPs was carried out with real-time PCR assays. Significant differences were observed between patients with DR and controls in the duration of diabetes (p < 0.001), insulin therapy (p < 0.001), glycated hemoglobin (p = 0.001), body mass index (p = 0.002), total cholesterol (p = 0.002), and low-density lipoprotein cholesterol (p < 0.001). However, we did not observe significant differences in the genotype and allele distribution of the two SNPs, between DR and control group (p < 0.05). Logistic regression analysis showed that rs6921438 and rs10738760 were not independent genetic risk factors for DR in the co-dominant model adjusted for the above-mentioned clinical and laboratory data. In conclusion, VEGF-related SNPs rs10738760 and rs6921438 are not associated with DR in our group of Slovenian patients (Caucasians) with T2DM.
Citations
Downloads
References
Pavljasević S, Pranjić N, Sarajlić D. Argon laser photo-coagulation complications in diabetic retinopathy. Bosn J Basic Med Sci 2004;4(2):41-4.
Kulenovic I, Rasic S, Karcic S. Development of microvascular complications in type 1 diabetic patients 10 years follow-up. Bosn J Basic Med Sci 2006;6(2):47-50.
Cekić S, Cvetković T, Jovanović I, Jovanović P, Pesić M, Stanković Babić G, et al. C-reactive protein and chitinase 3-like protein 1 as biomarkers of spatial redistribution of retinal blood vessels on digital retinal photography in patients with diabetic retinopathy. Bosn J Basic Med Sci 2014;14(3):177-84. https://doi.org/10.17305/bjbms.2014.3.21.
Petrovic MG, Kunej T, Peterlin B, Dovc P, Petrovic D. Gly482Ser polymorphism of the peroxisome proliferator-activated receptor-gamma coactivator-1 gene might be a risk factor for diabetic retinopathy in Slovene population (Caucasians) with type 2 diabetes and the Pro12Ala polymorphism of the PPARgamma gene is not. Diabetes Metab Res Rev 2005;21(5):470-4. https://doi.org/10.1002/dmrr.546.
Petrovic MG, Osredkar J, Saraga-Babić M, Petrovic D. K469E polymorphism of the intracellular adhesion molecule 1 gene is associated with proliferative diabetic retinopathy in Caucasians with type 2 diabetes. Clin Exp Ophthalmol 2008;36(5):468-72. https://doi.org/10.1111/j.1442-9071.2008.01785.x.
Nikolajević-Starčević J, Globočnik Petrovič M, Petrovič D. A1/A2 polymorphism of the glycoprotein IIIa gene and diabetic retinopathy in Caucasians with type 2 diabetes. Clin Exp Ophthalmol 2011;39(7):665-72. https://doi.org/10.1111/j.1442-9071.2011.02520.x.
Urbančič M, Kloboves, Prevodnik V, Petrovič D, Globočnik Petrovič M. A flow cytometric analysis of vitreous inflammatory cells in patients with proliferative diabetic retinopathy. Biomed Res Int 2013;2013:251528. https://doi.org/10.1155/2013/251528.
Petrovic MG, Kruzliak P, Petrovic D. The rs6060566 of the reactive oxygen species modulator 1 (Romo-1) gene affects Romo-1 expression and the development of diabetic retinopathy in Caucasians with type 2 diabetes. Acta Ophthalmol 2015;93(8):654-7. https://doi.org/10.1111/aos.12723.
Awata T, Inoue K, Kurihara S, Ohkubo T, Watanabe M, Inukai K, et al. A common polymorphism in the 5'-untranslated region of the VEGF gene is associated with diabetic retinopathy in type 2 diabetes. Diabetes 2002;51(5):1635-9. https://doi.org/10.2337/diabetes.51.5.1635.
Mankoč Ramuš S, Kumše T, Globočnik Petrovič M, Petrovič D, Cilenšek I. SNP rs2073618 of the osteoprotegerin gene is associated with diabetic retinopathy in Slovenian patients with type 2 diabetes. Biomed Res Int 2013;2013:364073. https://doi.org/10.1155/2013/364073.
Bates DO, Harper SJ. Regulation of vascular permeability by vascular endothelial growth factors. Vascul Pharmacol 2002;39(4-5):225-37.
https://doi.org/10.1016/S1537-1891(03)00011-9.
Jiang Y, Sun L, Xuan X, Wang J. Impacts of N-Butylphthalide on expression of growth factors in rats with focal cerebral ischemia. Bosn J Basic Med Sci 2016;16(2):102-7. https://doi.org/10.17305/bjbms.2016.560.
Wirostko B, Wong TY, Simó R. Vascular endothelial growth factor and diabetic complications. Prog Retin Eye Res 2008;27(6):608-21. https://doi.org/10.1016/j.preteyeres.2008.09.002.
Bonnefond A, Saulnier PJ, Stathopoulou MG, Grarup N, Ndiaye NC, Roussel R, et al. What is the contribution of two genetic variants regulating VEGF levels to type 2 diabetes risk and to microvascular complications? PLoS One 2013;8(2):e55921. https://doi.org/10.1371/journal.pone.0055921.
Berrahmoune H, Herbeth B, Lamont JV, Masson C, Fitzgerald PS, Visvikis-Siest S. Heritability for plasma VEGF concentration in the Stanislas family study. Ann Hum Genet 2007;71(Pt 1):54-63. https://doi.org/10.1111/j.1469-1809.2006.00298.x.
Globočnik Petrovič M, Korošec P, Košnik M, Osredkar J, Hawlina M, Peterlin B, et al. Local and genetic determinants of vascular endothelial growth factor expression in advanced proliferative diabetic retinopathy. Mol Vis 2008;14:1382-7.
Debette S, Visvikis-Siest S, Chen MH, Ndiaye NC, Song C, Destefano A, et al. Identification of cis- and trans-acting genetic variants explaining up to half the variation in circulating vascular endothelial growth factor levels. Circ Res 2011;109(5):554-63. https://doi.org/10.1161/CIRCRESAHA.111.243790.
Genuth S, Alberti KG, Bennett P, Buse J, Defronzo R, Kahn R, et al. Follow-up report on the diagnosis of diabetes mellitus. Diabetes Care 2003;26(11):3160-7. https://doi.org/10.2337/diacare.26.11.3160.
[No authors listed]. Grading diabetic retinopathy from stereoscopic color fundus photographs - an extension of the modified Airlie House classification. ETDRS report number 10. Early Treatment Diabetic Retinopathy Study Research Group. Ophthalmology 1991;98(5 Suppl):786-806. https://doi.org/10.1016/S0161-6420(13)38012-9.
Petrovic MG, Peterlin B, Hawlina M, Petrovic D. Aldose reductase (AC)n gene polymorphism and susceptibility to diabetic retinopathy in Type 2 diabetes in Caucasians. J Diabetes Complications 2005;19(2):70-3. https://doi.org/10.1016/j.jdiacomp.2004.08.004.
Petrovič D. Candidate genes for proliferative diabetic retinopathy. Biomed Res Int 2013;2013:540416. https://doi.org/10.1155/2013/540416.
Hampton BM, Schwartz SG, Brantley MA Jr, Flynn HW Jr. Update on genetics and diabetic retinopathy. Clin Ophthalmol 2015;9:2175-93. DOI: 10.2147/OPTH.S94508.
Turnpenny P, Ellard S. Emery's Elements of Medical Genetics. 15th ed. Exeter, UK: Elsevier; 2017.
Nakayama M, Yasue H, Yoshimura M, Shimasaki Y, Kugiyama K, Ogawa H, et al. T-786C mutation in the 5'-flanking region of the endothelial nitric oxide synthase gene is associated with coronary spasm. Circulation 1999;99(22):2864-70. https://doi.org/10.1161/01.CIR.99.22.2864.
Additional Files
Published
Issue
Section
Categories
License
Copyright (c) 2017 Bosnian Journal of Basic Medical Sciences

This work is licensed under a Creative Commons Attribution 4.0 International License.
How to Cite
Accepted 2017-08-09
Published 2017-11-20









