Insight into the emerging role of SARS-CoV-2 nonstructural and accessory proteins in modulation of multiple mechanisms of host innate defense
DOI:
https://doi.org/10.17305/bjbms.2020.5543Keywords:
COVID-19, SARS-CoV-2, nonstructural proteins, accessory proteins, proinflammatory, interferon, immune evasion, immunomodulatorsAbstract
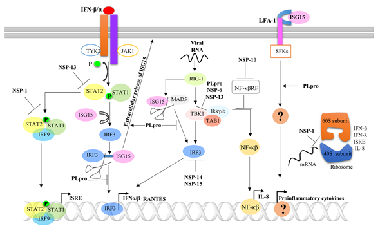
Coronavirus disease-19 (COVID-19) is an extremely infectious disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that has become a major global health concern. The induction of a coordinated immune response is crucial to the elimination of any pathogenic infection. However, SARS-CoV-2 can modulate the host immune system to favor viral adaptation and persistence within the host. The virus can counteract type I interferon (IFN-I) production, attenuating IFN-I signaling pathway activation and disrupting antigen presentation. Simultaneously, SARS-CoV-2 infection can enhance apoptosis and the production of inflammatory mediators, which ultimately results in increased disease severity. SARS-CoV-2 produces an array of effector molecules, including nonstructural proteins (NSPs) and open-reading frames (ORFs) accessory proteins. We describe the complex molecular interplay of SARS-CoV-2 NSPs and accessory proteins with the host’s signaling mediating immune evasion in the current review. In addition, the crucial role played by immunomodulation therapy to address immune evasion is discussed. Thus, the current review can provide new directions for the development of vaccines and specific therapies.
Citations
Downloads
Downloads
Additional Files
Published
Issue
Section
Categories
How to Cite
Accepted 2021-03-04
Published 2021-10-01









