Comprehensive analysis of a NAD+ metabolism-derived gene signature to predict the prognosis and immune landscape in endometrial cancer
DOI:
https://doi.org/10.17305/bb.2023.9489Keywords:
Nicotinamide adenine dinucleotide (NAD ), endometrial cancer (EC), prognostic signature, biomarker, immunotherapyAbstract
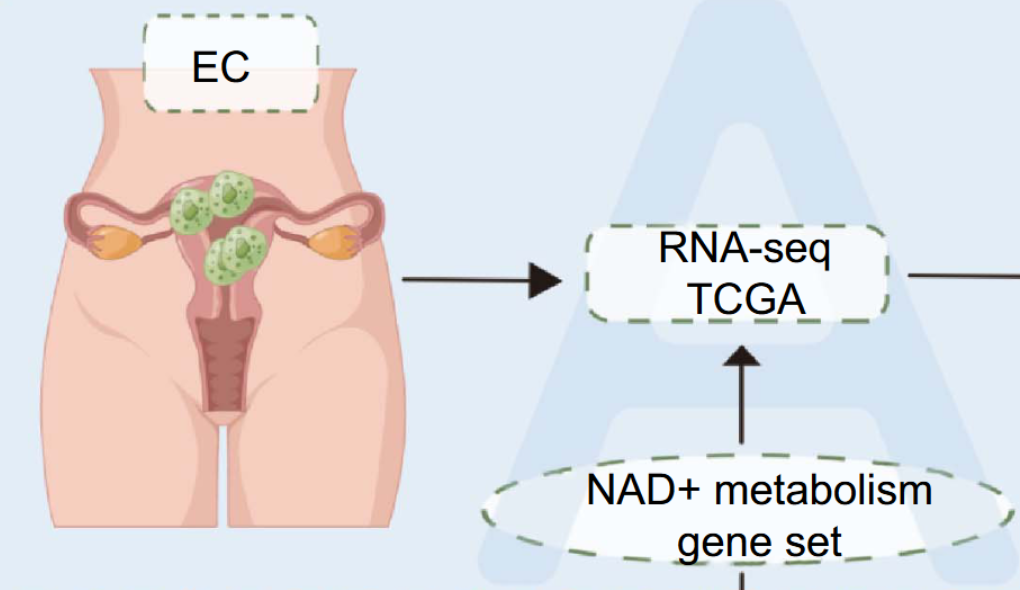
As a crucial regulator influencing tumor progression, nicotinamide adenine dinucleotide (NAD+) is widely acknowledged. However, its role in endometrial cancer (EC) is not completely understood. In this study, we aimed to develop an NAD+metabolic-related genes (NMRGs) risk signature that could reflect the prognosis of EC patients and their responsiveness to immunotherapy and chemotherapy. Data from The Cancer Genome Atlas (TCGA) databases and the Molecular Signatures Database (MSigDB) confirmed two distinct NMRG subtypes in EC patients using consensus clustering, and a risk score was constructed utilizing an NAD+-related prognostic signature depending on the least absolute shrinkage and selection operator (LASSO) Cox regression analysis. Receiver operating characteristic (ROC) curves were employed to assess the model’s precision. Additionally, we used Gene Set Enrichment Analysis (GSEA) to predict the biological signaling pathways that might be involved. We also explored the role of the risk score in immune cell infiltration, tumor mutation burden (TMB), immunotherapy, and chemotherapy. Our study established a prognostic risk signature based on six NMRGs, and we observed that the high-risk group was associated with a poorer prognosis. Furthermore, we identified a strong correlation between the high-risk group and several pathways, including DNA replication, cell cycle, and mismatch repair. Lastly, our findings highlighted the influence of NMRGs on the regulation of immune infiltration in EC. Therefore, this signature holds potential value in predicting the prognosis of EC patients and guiding their management, including decisions regarding immunotherapy and chemotherapy, ultimately improving the accuracy of EC patient care.
Citations
Downloads
Downloads
Additional Files
Published
Issue
Section
Categories
License
Copyright (c) 2023 Dan Hu, JunHong Du, YueMei Cheng, YiJuan Xing, RuiFen He, Xiaolei Liang, HongLi Li, YongXiu Yang

This work is licensed under a Creative Commons Attribution 4.0 International License.
How to Cite
Accepted 2023-08-29
Published 2024-03-11









